Every cell is a master builder, able to craft useful and structurally complex molecules, time and again and with astonishingly few mistakes. Scientists are keen to replicate this feat to build their own molecular factories, but first, they’ll need to understand it.

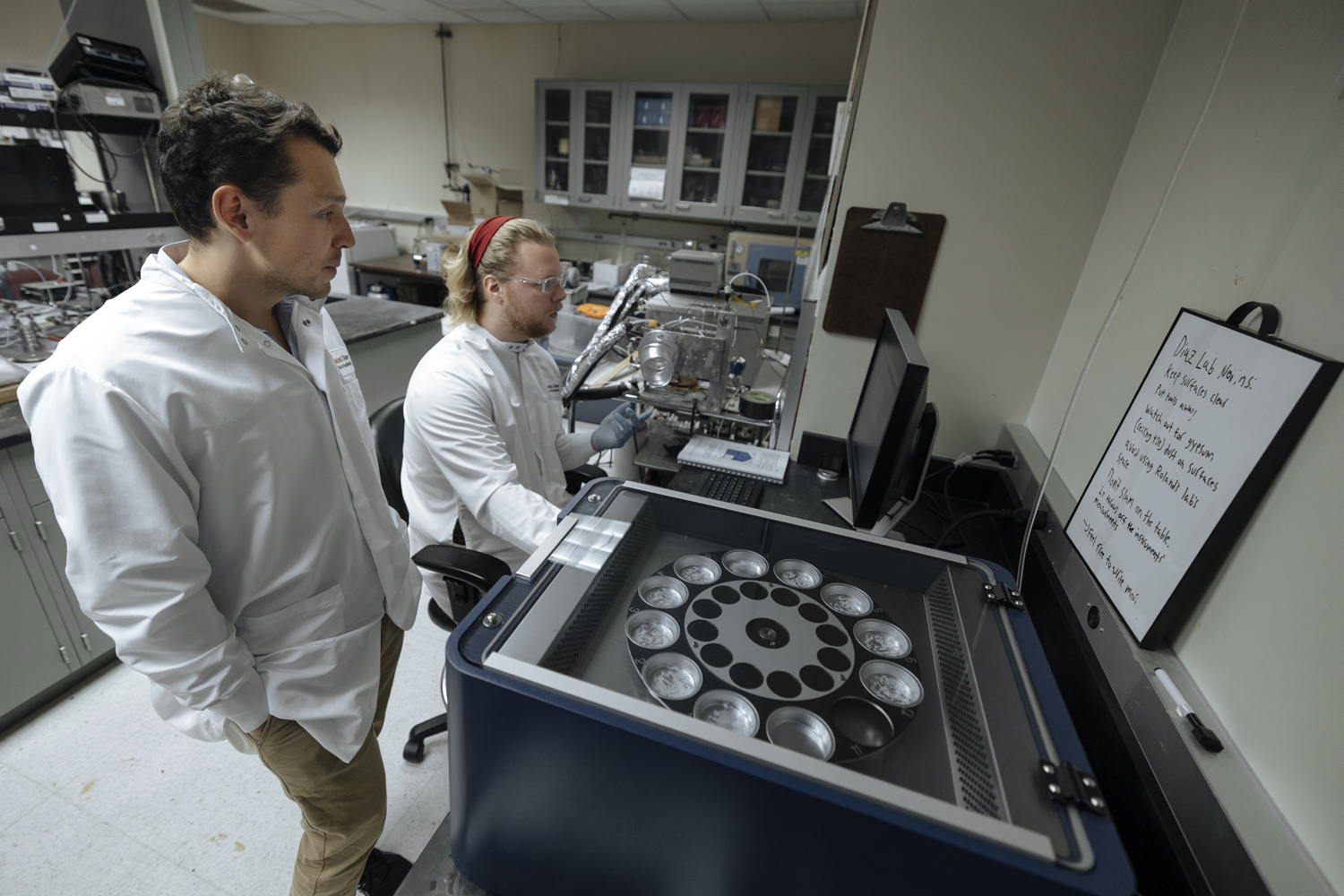
Researchers investigate how molecular assembly lines maintain their precise control while shepherding growing molecules through a complex, multi-step construction process. (Image credit: Getty Images)
“We have thousands of these assembly lines in nature, and they all make unique compounds,” said Dillon Cogan, a postdoctoral scholar in the lab of Chaitan Khosla, a professor of chemistry and chemical engineering at Stanford. “The dream is to one day be able to recombine pieces from different assembly lines so that we can make useful compounds not found in nature. To do that, we need to know the design principles that make these things work.”
In a study published Nov. 5 in Science, researchers from Stanford University use one of the most sophisticated structural biology techniques available to investigate how these molecular assembly lines maintain their precise control while shepherding growing molecules through a complex, multi-step construction process.
The molecules in question are called polyketides, a category that includes drugs and antibiotics. Cells synthesize polyketides through molecular assembly lines called synthases.
Each synthase contains anywhere between three to 30 “modules,” groups of active proteins, or enzymes, organized sequentially. Each module is a station in the assembly line that is responsible for adding a piece to a growing molecular chain and then installing chemical modifications to that unit. Passing from module to module, a polyketide grows in size and complexity until it eventually rolls off the conveyor belt in its final form.
Khosla, Cogan and colleagues focused on a module from the assembly line that produces the antibiotic erythromycin. They wanted to understand how this assembly line, like others, always manages to push the growing molecule in the right direction, a feat that the laws of thermodynamics can’t fully explain.
To do so, they turned to Wah Chiu, a professor at Stanford and at the SLAC National Accelerator Laboratory, which Stanford operates for the U.S. Department of Energy (DOE). Chiu is an expert in a sophisticated technique called cryogenic electron microscopy (cryo-EM), which captures multiple snapshots of moving proteins in slightly different configurations. Applying cryo-EM to the assembly line module would allow researchers to observe it in various shapes, each one corresponding to a different stage in the assembly line process, which is something other techniques hadn’t revealed.
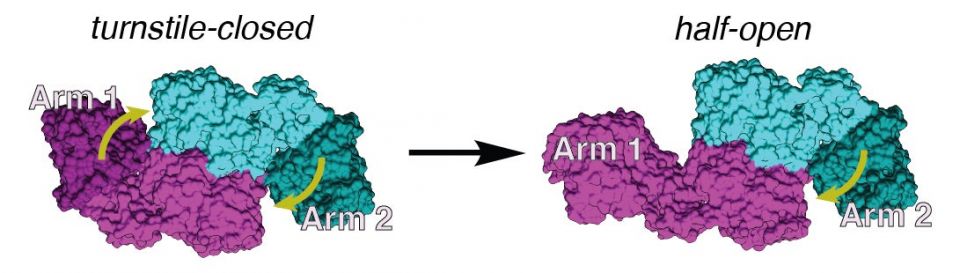
An image, based on cryo-EM data, shows the difference between the symmetric, turnstile-closed conformation of the assembly line module (left) and the asymmetric, bent-arm conformation (right). The pair of identical enzymes that make up the module are shown in cyan and magenta. When the two arms are closed (left), incoming molecules are blocked from entering (represented by yellow arrows). When one arm is bent (right), it allows a single molecule to enter the module. (Image credit: Dillon Cogan)
“They showed us what the assembly line looks like when it’s not processing molecules. It’s like a Sunday at the BMW plant. We wanted to see the plant on a Monday,” said Khosla, who is the Wells H. Rauser and Harold M. Petiprin Professor of Chemical Engineering and Chemistry at Stanford.
Chiu, the Wallenberg-Bienenstock Professor at Stanford and a Professor of Photon Science at SLAC, was instantly intrigued. “These are amazingly complex molecular machines. There are so many components that have to come together at the right place and the right time, in a highly orchestrated way, for them to work,” said Chiu.
Chiu asked Kaiming Zhang, a former postdoctoral scholar in his lab, to partner with Cogan to study the assembly line module at the Stanford-SLAC Cryo-Electron Microscopy Facility.
After many years of work, the pair glimpsed something unexpected. Each module is made up of a pair of enzymes, each of which has a molecular arm that extends out from the module’s sides. It was widely thought that these arms mirror one another in their poses. But in the module Zhang and Cogan examined, one arm extended out while the second arm flexed downward.
The pair realized that the structure they were observing was actually the module in action and that the bent arm could be the key to the assembly line’s directionality.
The finding also helped resolve another mystery that Khosla’s lab was grappling with. His group had previously found that each module can only work on two molecules at a time. They called this a “turnstile” mechanism, with each module closing itself off to incoming chains until it releases one it’s working on. But what they didn’t know was how it closed itself off. Now, they think this flexed arm acts like the arm of the turnstile.
The turnstile arm appears to have two jobs. First, it acts as a gatekeeper and physically blocks incoming molecules from entering while one is being processed. Second, the contortion of the enzyme into that asymmetric pose requires energy, which gets stored in the flex of the arm. The team hypothesized that the relaxation of the arm back to its “normal” state, which releases the pent-up energy, might help propel the molecule under construction to the next stage of the assembly line.
“That these enzymes are capturing energy in these amazing contortions, and that they use that energy to power something else – in this case, direction – is so exciting,” said Khosla.
The team’s hypothesis that asymmetry helps confer directional selectivity to the assembly lines is supported by a second paper, published in the same issue of Science. Scientists at the University of Texas at El Paso, Cornell University and SLAC used both cryo-EM and a technique called X-ray crystallography to study a module from a different polyketide synthase and observed a similar asymmetric, flexed-arm conformation.
“That both our groups were able to tackle this complex biological system is a testament to the huge investment Stanford and SLAC have made in structural biology infrastructure,” said Khosla.
Khosla is an Institute Scholar at Stanford ChEM-H; the director of the Innovative Medicines Accelerator; and a member of Bio-X, the Maternal & Child Health Research Institute, the Stanford Cancer Institute and the Wu Tsai Neurosciences Institute. Chiu is also a Faculty Fellow at Stanford ChEM-H, and a member of Bio-X, the Cardiovascular Institute, and the Wu Tsai Neurosciences Institute. Zhang is now at the University of Science and Technology of China. Other co-authors include former graduate student Xiuyuan Li; former postdoc Shanshan Li, now of the University of Science and Technology of China; Grigore Pintilie, Research Scientist at Stanford; former postdoc Soung-Hun Roh, now of Seoul National University; and Charles Craik of the University of California San Francisco.
The research was supported by the National Institutes of Health.
To read all stories about Stanford science, subscribe to the biweekly Stanford Science Digest.
Media Contacts
Rebecca McClellan, Stanford ChEM-H: rmcclell@stanford.edu
Author
Rebecca McClellan





