
Postdoctoral scholar Laurent Formery (left) and biology Professor Christopher Lowe with starfish on the shore of Stanford’s Hopkins Marine Station, in Monterey, California. (Image credit: LiPo Ching / Stanford University)
If you put a hat on a starfish, where would you put it? On the center of the starfish? Or on the point of an arm and, if so, which one? The question is silly, but it gets at serious questions in the fields of zoology and developmental biology that have perplexed veteran scientists and schoolchildren in introductory biology classes alike: Where is the head on a starfish? And how does their body layout relate to ours?
Now, a new Stanford study that used genetic and molecular tools to map out the body regions of starfish – by creating a 3D atlas of their gene expression – helps answer this longstanding mystery. The “head” of a starfish, the researchers found, is not in any one place. Instead, the headlike regions are distributed with some in the center of the sea star as well as in the center of each limb of its body.
“The answer is much more complicated than we expected,” said Laurent Formery, lead author and postdoc in the labs of Christopher Lowe at the Stanford School of Humanities and Sciences and Daniel S. Rokhsar at the University of California, Berkeley. “It is just weird, and most likely the evolution of the group was even more complicated than this.”
Starfish (sea stars) belong to a group of animals called echinoderms. Echinoderms and humans are closely related, yet the life cycle and anatomy of sea stars are very different from ours.
Sea stars begin life as fertilized eggs that hatch into a free-floating larva. The larvae bob in the ocean in a plankton form for weeks to months before settling to the ocean floor to perform a magic trick of sorts – transforming from a bilateral (symmetric across the midline) body plan into an adult with a five-point star shape called a pentaradial body plan.
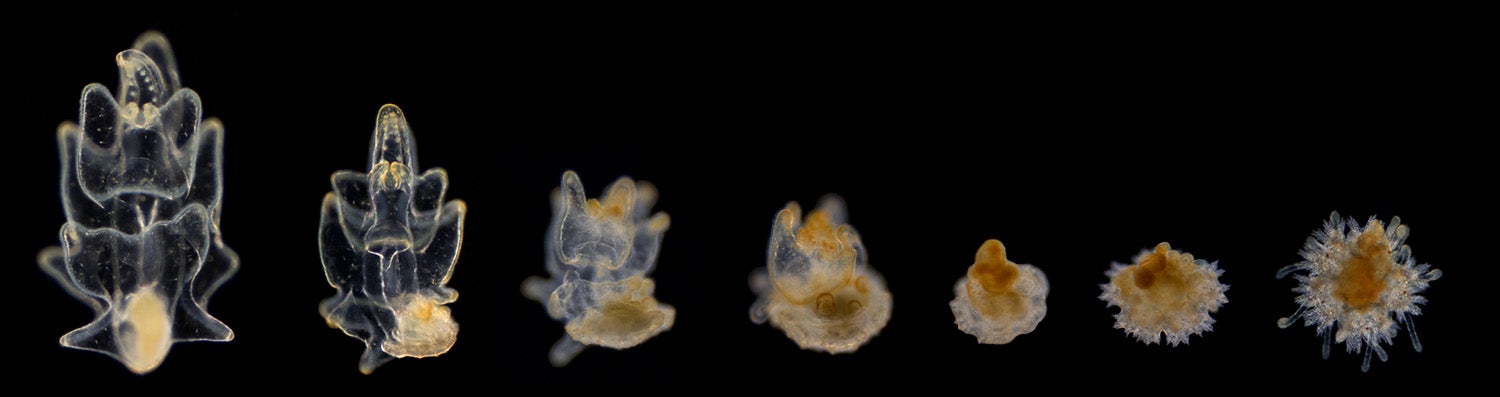
Images of sea star metamorphosis, where they transition from larvae with a bilateral (symmetric across the midline) body plan into young adult sea stars with a five-point star shape called a pentaradial body plan. (Image credit: Laurent Formery)
“This has been a zoological mystery for centuries,” said Lowe, who is also a researcher at Hopkins Marine Station and senior author of the paper that published Nov. 1 in Nature. “How can you go from a bilateral body plan to a pentaradial plan, and how can you compare any part of the starfish to our own body plan?”
Mapping stars
For puzzles such as this one, researchers often conduct comparative studies to identify similar structures in related groups of animals to glean clues about the evolutionary events that prompted the trait of interest.
“The problem with starfish is there is nothing on a starfish anatomically that you can relate to a vertebrate,” said Lowe. “There is just nothing there.”
At least, nothing on the outside of a starfish. And that is where genetic and molecular techniques come in.
During his graduate research, Formery studied early development in sea urchins – echinoderms, like sea stars, that also start their life as bilateral larvae before transforming into adults with fivefold symmetry. When Formery joined Lowe’s lab, Formery’s knowledge of echinoderm development combined with Lowe’s expertise in molecular biology techniques to help tackle the mystery of sea stars’ baffling body plan.
The team used a group of well-studied molecular markers (Hox genes are an example) that act as blueprints for an organism’s body plan by “telling” each cell which body region it belongs to.
“If you strip away the skin of an animal and look at the genes involved in defining a head from a tail, the same genes code for these body regions across all groups of animals,” said Lowe. “So we ignored the anatomy and asked: Is there a molecular axis hidden under all this weird anatomy and what is its role in a starfish forming a pentaradial body plan?”
To investigate this question, the researchers used RNA tomography, a technique that pinpoints where genes are expressed in tissue, and in situ hybridization, a technique that zeroes in on a specific RNA sequence in a cell.
“First we sectioned sea star arms into thin slices from tip to center, top to bottom, and left to right,” said Formery, noting that sea stars regenerate missing limbs. “We used RNA tomography to determine which genes were expressed in each slice and then ‘reassembled’ the slices using computer models. This gave us a 3D map of gene expression.”
“In the second method, in situ hybridization chain reaction, we stained sea star tissue and visually inspected the samples to see where a gene was expressed,” said Formery. This enabled the researchers to examine anterior-posterior (head to tail) body patterning in the outermost layer of cells called the ectoderm.
“This was made possible by the recent, big, technical improvement in in situ hybridization, known as in situ hybridization chain reaction, Formery said. “This new method provides better resolution of where the gene is expressed.”
The research revealed that sea stars have a headlike territory in the center of each “arm” and a tail-like region along the perimeter. In an unexpected twist, no part of the sea star ectoderm expresses a “trunk” genetic patterning program, suggesting that sea stars are mostly headlike.
Mining truly diverse biodiversity
Research is often centered on groups of animals that look like us, the researchers explained. But if we focus on the familiar, we are less likely to learn something new.
“There are 34 different animal phyla living on this planet and in over roughly 600 million years they have all come up with different solutions to the same fundamental biological problems,” Lowe said. “Most animals don’t have spectacular nervous systems and are out chasing prey – they are modest animals that live in burrows in the ocean. People are generally not drawn to these animals, and yet they probably represent how much of life got started.”
This study demonstrates how a comparative approach that uses genetic and molecular techniques can be used to mine biodiversity for insights into why different animals look the way they do and how their body plans evolved.
“Even in recent molecular papers there’s a question mark near echinoderms on the evolutionary tree because we don’t know much about them,” Formery said. “It was nice to show that – at least at the molecular level – we have a new piece of the puzzle that can now be put on the tree.”
Formery, Lowe, and Rokhsar are also researchers at the Chan Zuckerberg Biohub San Francisco. Rokhsar is also a researcher at the Okinawa Institute of Science and Technology. Additional Stanford co-authors are Ian Kohnle, Judith Malnick, and Kevin Uhlinger of Hopkins Marine Station. Additional authors are from Pacific Biosciences in Menlo Park, California, and Columbia Equine Hospital in Gresham, Oregon.
This research was funded by NASA, the National Science Foundation, and the Chan Zuckerberg Biohub San Francisco.
Media Contacts
Holly Alyssa MacCormick, Stanford School of Humanities and Sciences: hollymac@stanford.edu
